NGS Analysis of Hematology-Oncology Clonality Testing
This news was updated in April 2024
Invivoscribe’s NGS LymphoTrack® Dx Assays are used to detect clonal gene rearrangements, somatic hypermutations and for the study of measurable residual disease (MRD). The full clonality suite of LymphoTrack® Dx Assays are CE-marked and developed for use with the leading NGS platforms, including optimized multiplex PCR master mixes with primers incorporating platform-specific adapters and specimen tracking sequencing identification tags for a one-step PCR workflow. A comprehensive bioinformatics software package is provided free of charge with purchase; enabling you to identify the DNA sequence, clonal prevalence, V-J family identity for each gene rearrangement, and with the IGHV Leader or IGH FR1 assay, the extent of IGHV somatic hypermutation (SHM).
Key benefits of LymphoTrack® Dx Assays
- Common sources of DNA – Formalin-fixed paraffin-embedded (FFPE), fresh/frozen tissue, peripheral blood, bone marrow
- One-Step PCR Master Mixes
- Multiplexing – Reduces Costs by Combining:
- Multiple Samples - up to 12 (Ion S5/PGM™) & 24 (MiSeq®)
- Multiple Invivoscribe Assays - up to 72 or 168 Samples
- Other Platform-Specific Assays
- Unparalleled Sensitivity
- Unparalleled clonality detection with the ability to identify and track the specific sequence of clonal populations for MRD research studies.
- Analysis Software Package - Included
Simple workflow
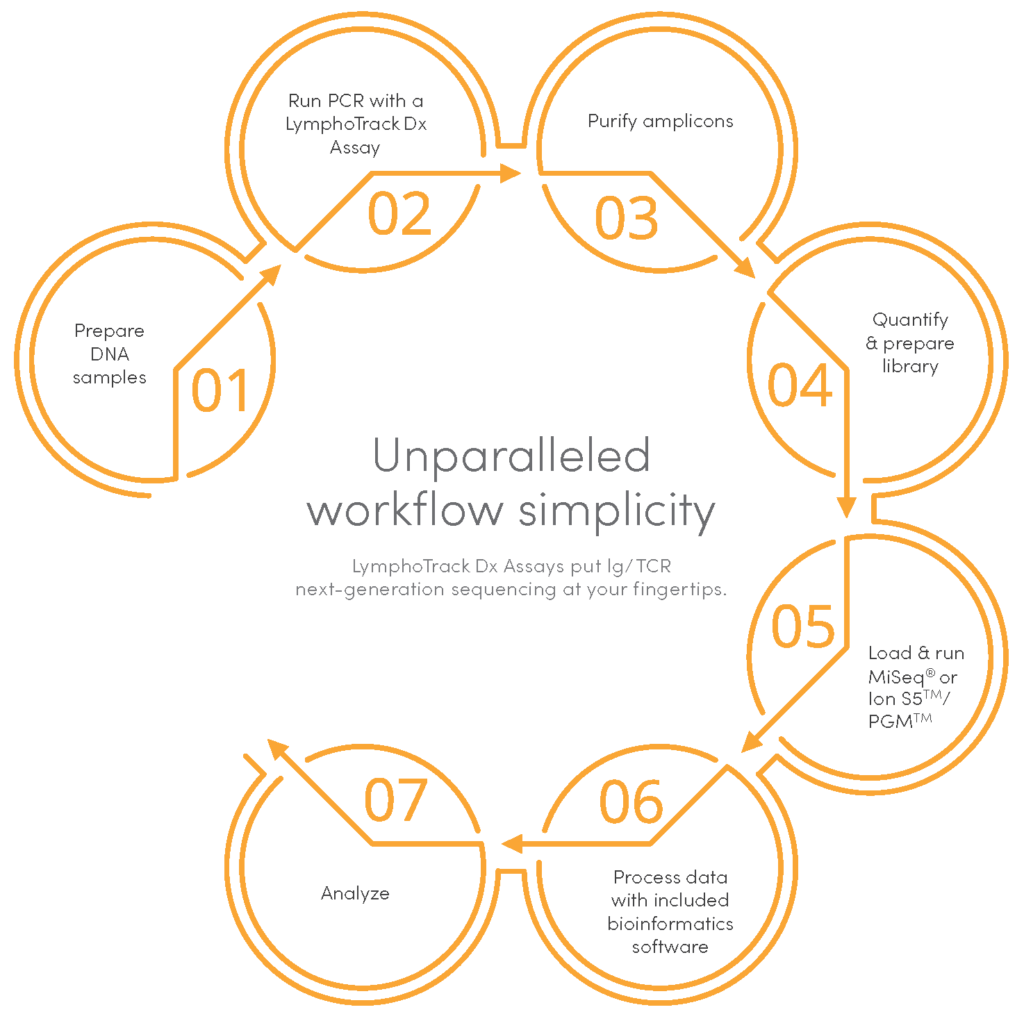
Multiplexing capabilities
- Kits include up to 24 indices to increase workflow flexibility
- Designed with the ability to pool multiple targets in a single sequencing run
- Decreased testing costs
- Increased throughput capabilities
LymphoTrack Dx software
- Included with each LymphoTrack® Dx kit
- Available for both Ion Torrent S5/PGM™ and Illumina® MiSeq® platforms
- Does not require bioinformatics personnel
- Can run on most standard Windows platforms
- Utilizes FASTQ files
- Fast results reported in individual PDF reports or in Excel
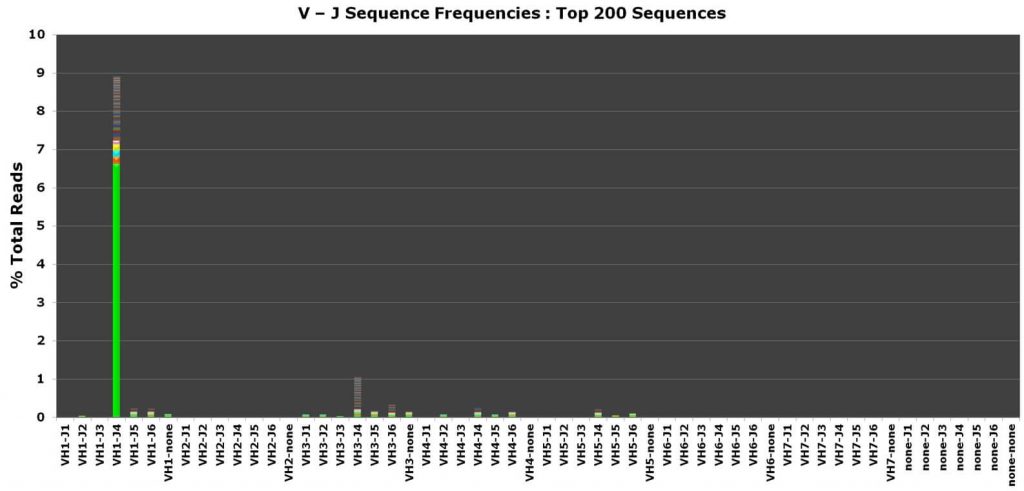
Data interpretation
- Easy clinical interpretation for the evidence of clonality
- DNA sequence, frequency, length, and family directly revealed
- Somatic Hypermutation status calculated from IGHV sequencing data
- Backed by clinical studies
- Clonal sequences can be tracked in subsequent samples
LymphoTrack® Dx Assays for the Illumina MiSeq®
- LymphoTrack Dx IGH FR1/2/3 Assay - MiSeq
- LymphoTrack Dx IGHV Leader Somatic Hypermutation Assay – MiSeq
- LymphoTrack Dx IGK Assay – MiSeq
- LymphoTrack Dx TRB Assay – MiSeq
- LymphoTrack Dx TRG Assay – MiSeq
LymphoTrack® Dx Assays for the Ion S5™/PGM™
- LymphoTrack Dx IGH FR1/2/3 Assay – S5/PGM
- LymphoTrack Dx IGK Assay – S5/PGM
- LymphoTrack Dx TRG Assay – S5/PGM
References
- Stamatopoulos B, Timbs A, Bruce D, et al. Targeted deep sequencing reveals clinically relevant subclonal IgHV rearrangements in chronic lymphocytic leukemia. Leukemia. 2017;31(4):837-845.
- Shin S, Hwang IS, Kim J, Lee KA, Lee ST, Choi JR. Detection of immunoglobulin heavy chain gene clonality by next-generation sequencing for minimal residual disease monitoring in b-lymphoblastic leukemia. Ann Lab Med. 2017;37(4):331-335.
- Lamarche C, Orio J, Georges-Tobar V, et al. Clinical-scale rapid autologous bk virus-specific t cell line generation from kidney transplant recipients with active viremia for adoptive immunotherapy. Transplantation. 2017;101(11):2713-2721.
- Levy-Mendelovich S, Lev A, Rechavi E, et al. T and B cell clonal expansion in Ras-associated lymphoproliferative disease (Rald) as revealed by next-generation sequencing. Clin Exp Immunol. 2017;189(3):310-317.
PCR | B- & T-cell clonality assays (IGH, IGK, TRG, TRB) | Chromosome Translocations | CDx FLT3 Mutation Assay | NGS Gene Panels | MRD clonality solutions | Molecular Biology | Services for Oncology Testing